Interactive Biophysical Signal Plotting
A dashboard designed for summer research interns in the lab who are unfamiliar with programming. It serves as an easy tool for visualizing data.
This dashboard is made with RShiny. You can access and interact with it here
The default option is to use the sample data, which is a sample experimental data obtained from my PhD work, containing torque and sEMG traces sampled at 1kHz. You can also upload your own experimental data, with the caveat noted below.
This dashboard is made specifically for preliminary data analyses in the Gurari Lab, so that uploaded data should follow the following structure: column 1 - time, column 2 - state, column 3 & 4 - torque / load cell signals, column 5:12 - eight sEMG signals.
It currently does not have ability to generalize to other data structures. Uploaded data not following the structure specified above may not get displayed correctly. However, you can view, fork, and modify my source code on my github repository, specifically the loadSensorData.R script to fit your data structure if needed.
This dashboard has two main functions:
- Plot and the elbow and shoulder torque traces and zoom in to see details.
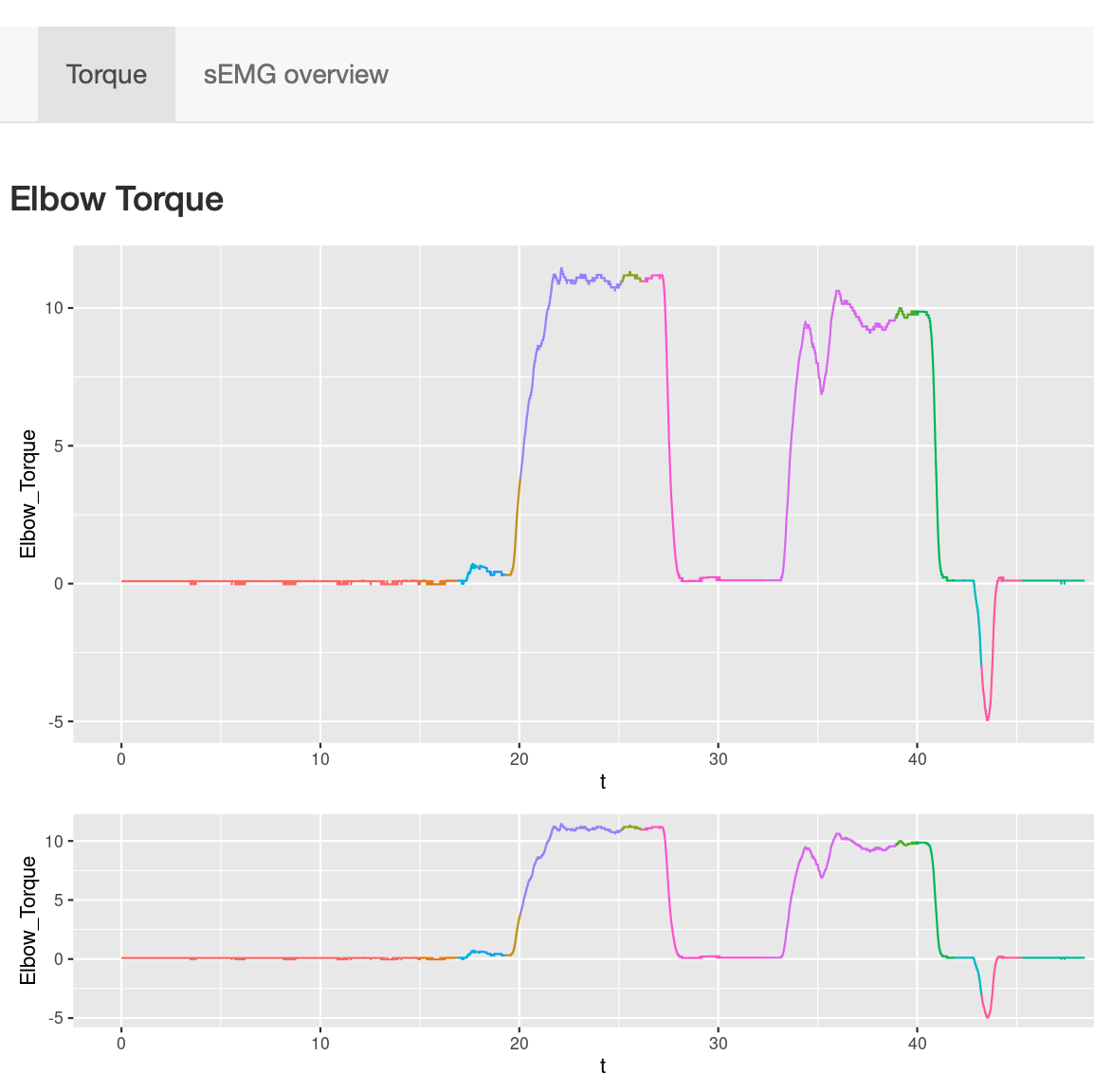
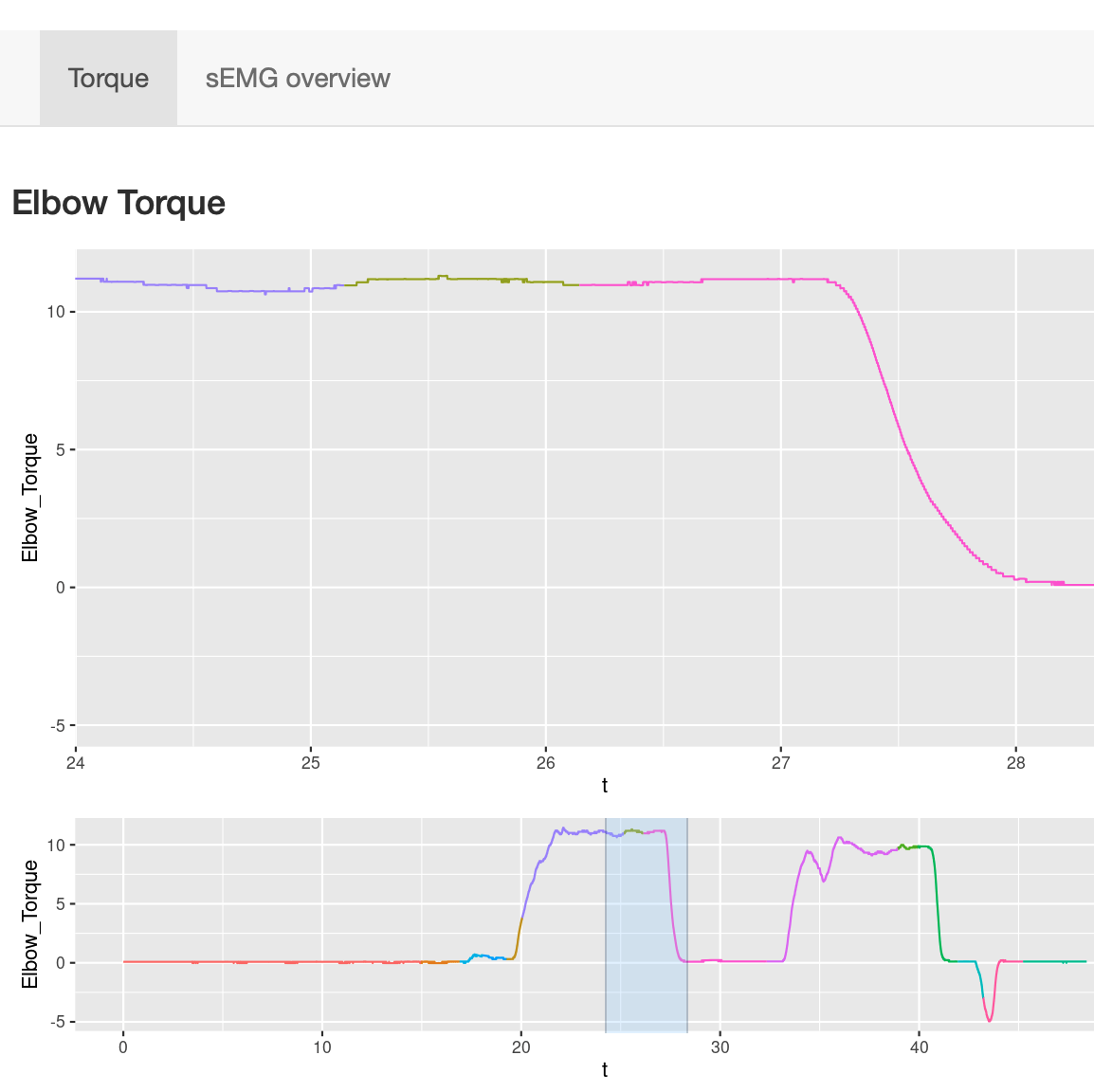 Users can select time frame of interest from the lower panel and zoom into the corresponding torque traces.
Users can select time frame of interest from the lower panel and zoom into the corresponding torque traces.This function is particularly useful for visually inspecting whether individuals maintained torques at any given time point of the experiment.
- Plot sEMG signals with synchronized zoom functions.
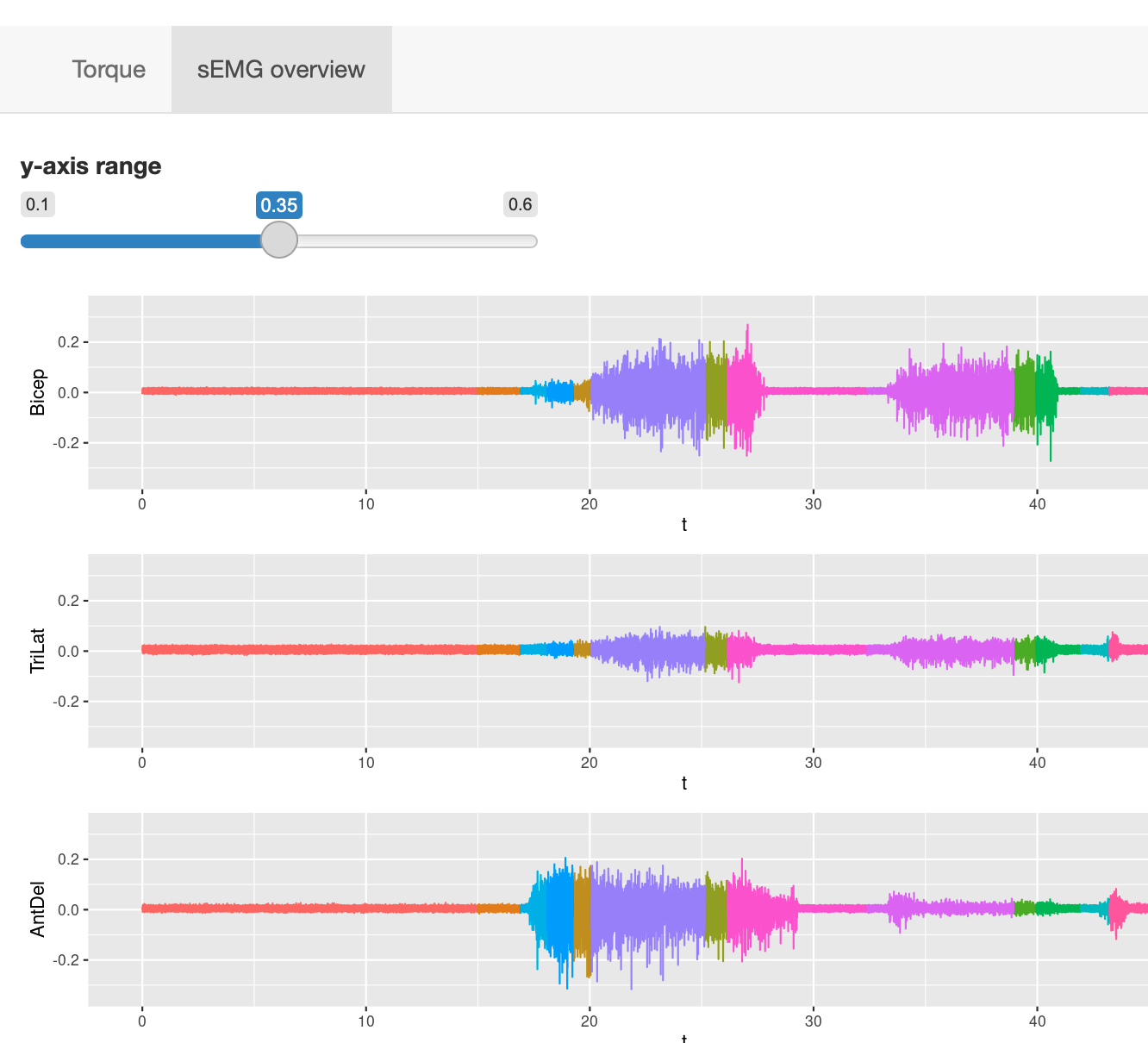
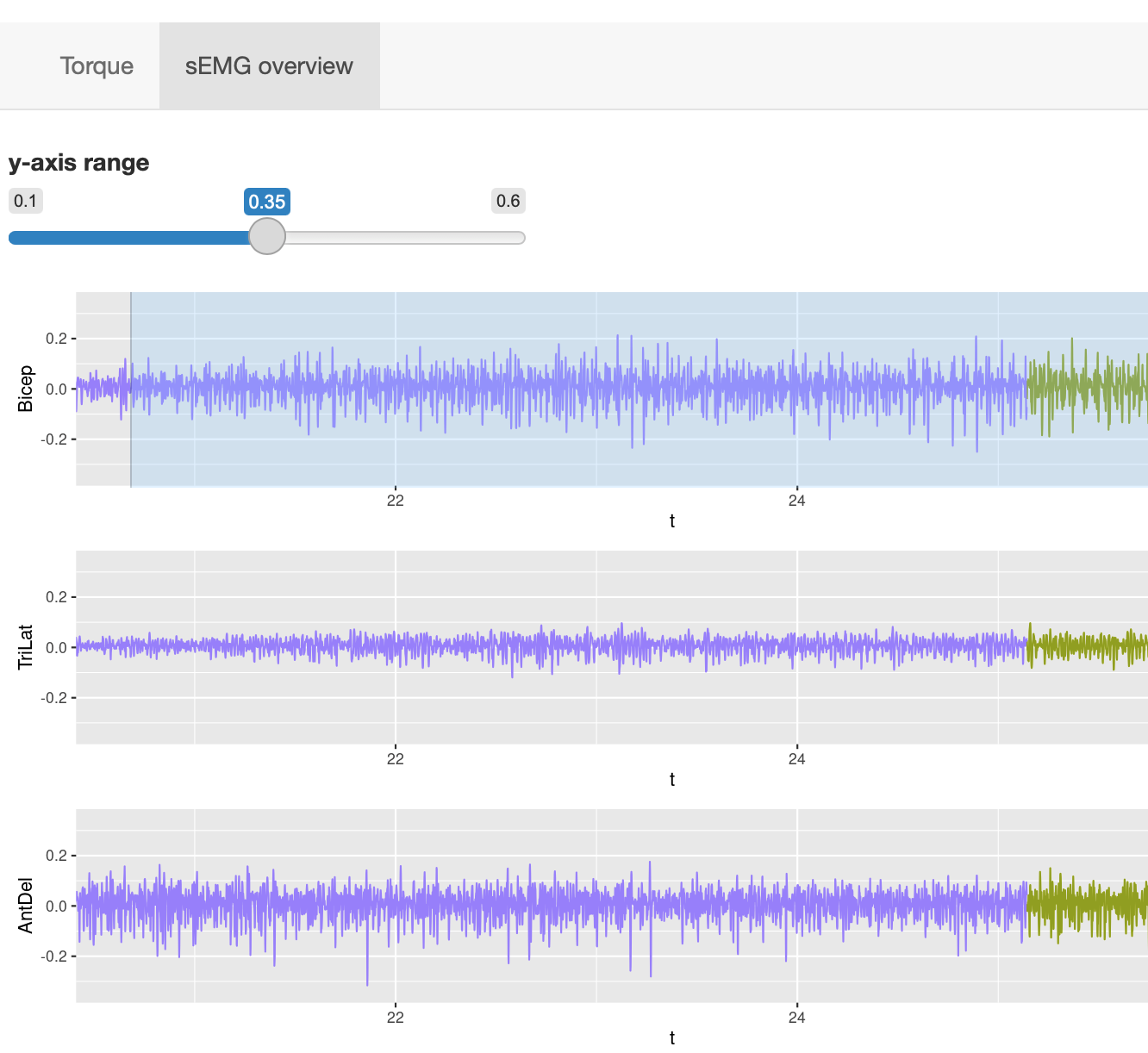 Users can zoom into specific time segments by selecting on any of the EMG traces, and adjust the y-axis limits.
Users can zoom into specific time segments by selecting on any of the EMG traces, and adjust the y-axis limits.This function is useful for visually inspecting all the EMG signal quality as well as observing the change in all eight muscles activiities at the same time.
More functions to come:
- Rectify button to rectify the EMG signals
- Filter button to apply different filters to smooth the EMG signals
- …